- Information
- Symbol: ERS2
- MSU: LOC_Os05g06320
- RAPdb: Os05g0155200
- PSP score
- LOC_Os05g06320.2: 0.0021
- LOC_Os05g06320.3: 0.0168
- PLAAC score
- LOC_Os05g06320.2: 0
- LOC_Os05g06320.3: 0
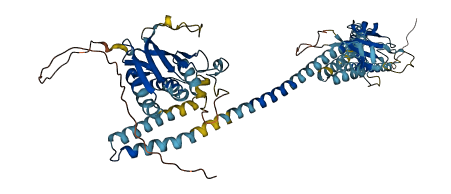
- pLDDT score
- 77.51
- Protein Structure from AlphaFold and UniProt
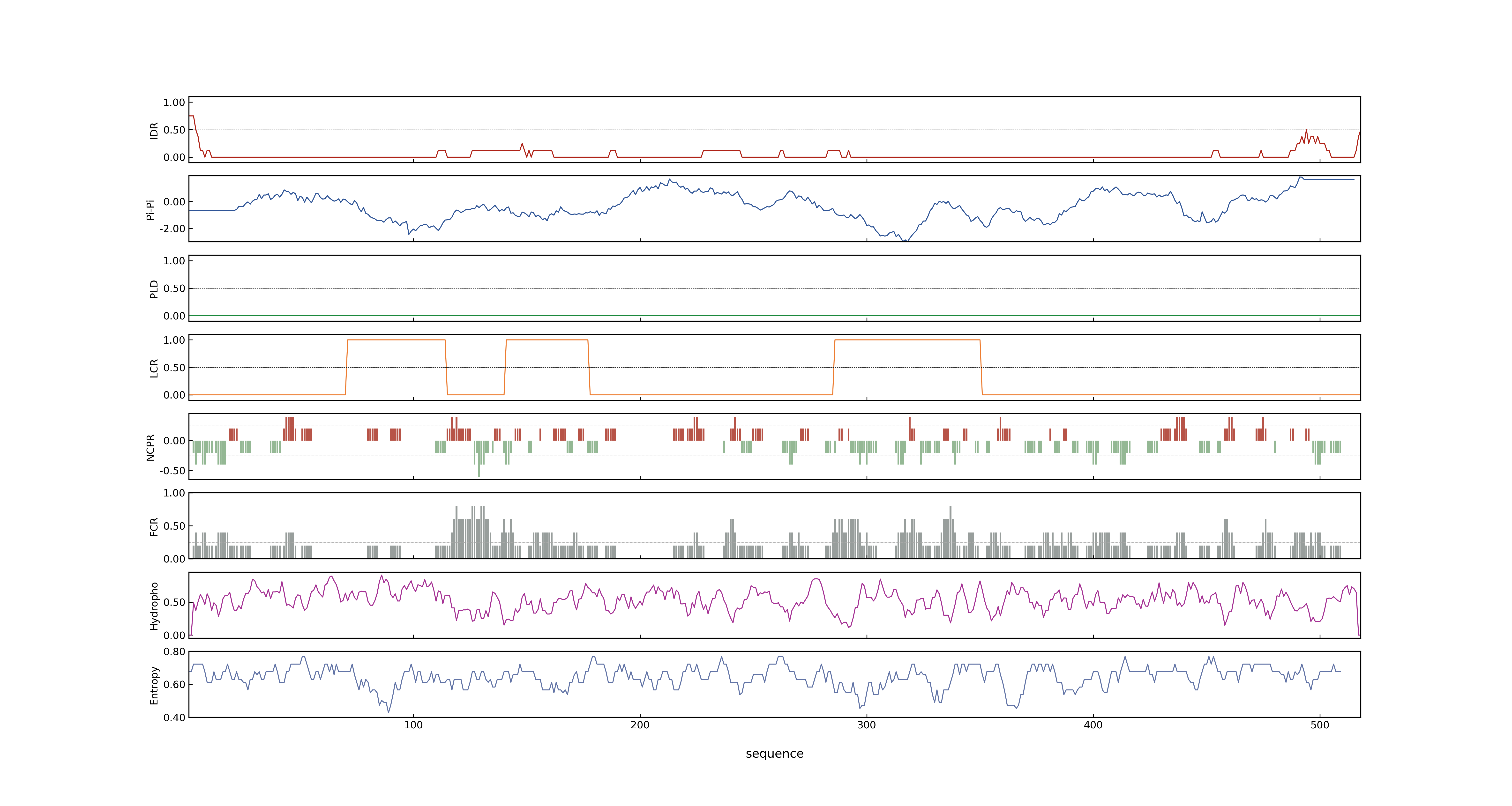
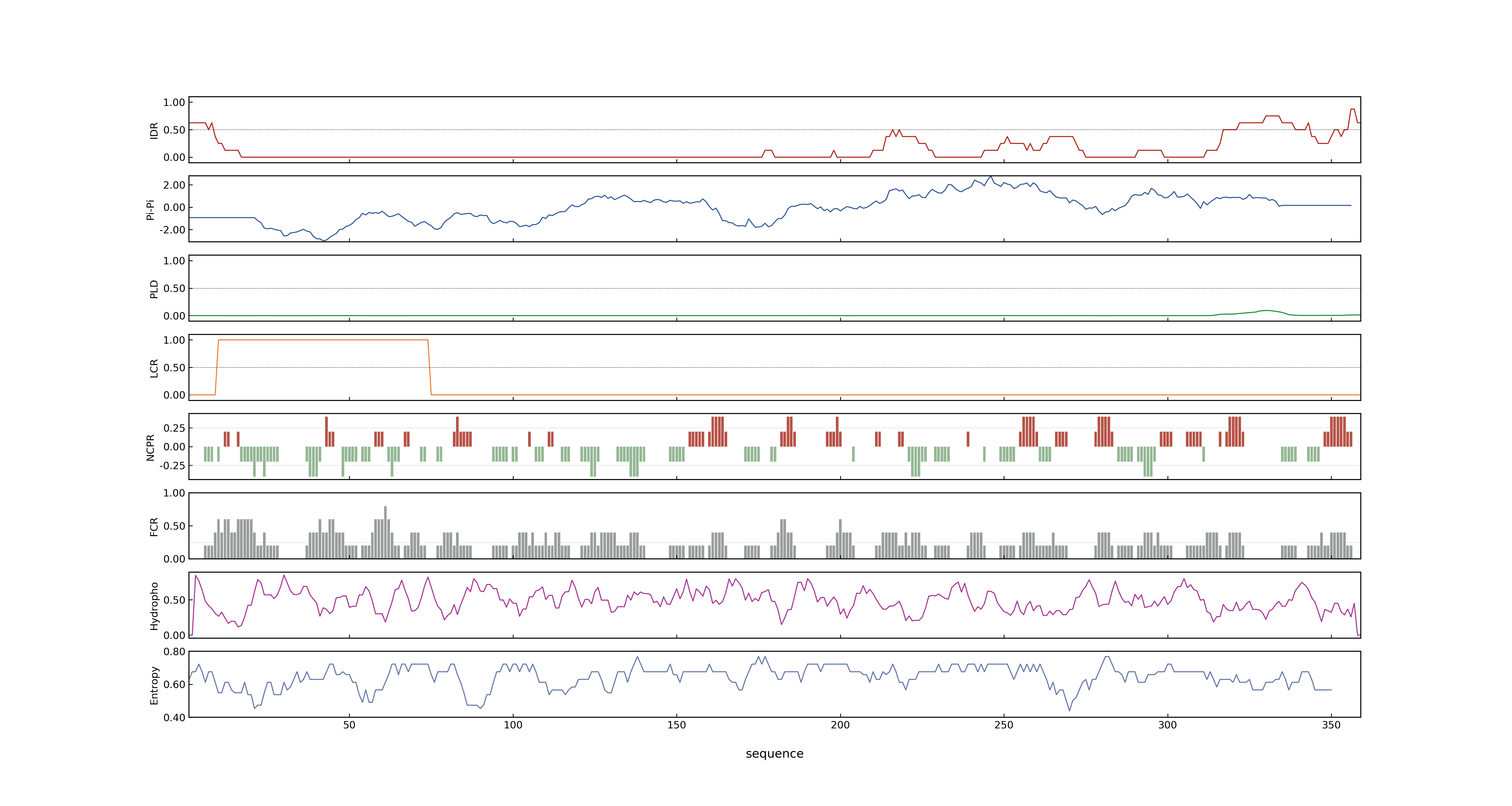
- MolPhase score
- LOC_Os05g06320.2: 0.31615442
- LOC_Os05g06320.3: 0.20527683
- MolPhase Result
- Publication
- Whole-genome analysis of Oryza sativa reveals similar architecture of two-component signaling machinery with Arabidopsis, 2006, Plant Physiol.
- Nomenclature for two-component signaling elements of rice, 2007, Plant Physiol.
- Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, 2004, J Exp Bot.
-
Genbank accession number
- Key message
- However, the abundance of OS-ERS2 mRNA was shown to be down-regulated by both IAA and ethylene treatments, indicating that it was not positively regulated by ethylene
- Five ethylene receptor genes, OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 were isolated and characterized from rice
- Deduced amino acid sequences of OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 showed that they exhibited significant homology to the prokaryotic two-component signal transducer and a wide range of ethylene receptors in a variety of plant species
- Analysis of the expression of the three ethylene receptor genes in different tissues of rice has unravelled their corresponding tissue-specificity in which OS-ERS1 was constitutively expressed in considerable amounts in all tissues studied, while OS-ERS2 and OS-ETR2 exhibited differential expression patterns in different tissues of rice
- Connection
- ERS1~OsERS1, ERS2, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, Five ethylene receptor genes, OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 were isolated and characterized from rice
- ERS1~OsERS1, ERS2, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, The genomic structure of OS-ERS1 and OS-ERS2 revealed that the introns within the coding sequences occurred in conserved positions to those of At-ETR1 and At-ERS1, whereas each of the OS-ETR2, OS-ETR3, and OS-ETR4 genes contained 1 intron within its coding region located at a position equivalent to those of At-ERS2, At-ETR2, and At-EIN4
- ERS1~OsERS1, ERS2, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, Deduced amino acid sequences of OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 showed that they exhibited significant homology to the prokaryotic two-component signal transducer and a wide range of ethylene receptors in a variety of plant species
- ERS1~OsERS1, ERS2, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, Analysis of the expression of the three ethylene receptor genes in different tissues of rice has unravelled their corresponding tissue-specificity in which OS-ERS1 was constitutively expressed in considerable amounts in all tissues studied, while OS-ERS2 and OS-ETR2 exhibited differential expression patterns in different tissues of rice
- ERS2, ETR3, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, Five ethylene receptor genes, OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 were isolated and characterized from rice
- ERS2, ETR3, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, The genomic structure of OS-ERS1 and OS-ERS2 revealed that the introns within the coding sequences occurred in conserved positions to those of At-ETR1 and At-ERS1, whereas each of the OS-ETR2, OS-ETR3, and OS-ETR4 genes contained 1 intron within its coding region located at a position equivalent to those of At-ERS2, At-ETR2, and At-EIN4
- ERS2, ETR3, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, Deduced amino acid sequences of OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 showed that they exhibited significant homology to the prokaryotic two-component signal transducer and a wide range of ethylene receptors in a variety of plant species
- ERS2, ETR4, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, Five ethylene receptor genes, OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 were isolated and characterized from rice
- ERS2, ETR4, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, The genomic structure of OS-ERS1 and OS-ERS2 revealed that the introns within the coding sequences occurred in conserved positions to those of At-ETR1 and At-ERS1, whereas each of the OS-ETR2, OS-ETR3, and OS-ETR4 genes contained 1 intron within its coding region located at a position equivalent to those of At-ERS2, At-ETR2, and At-EIN4
- ERS2, ETR4, Differential expression of three genes encoding an ethylene receptor in rice during development, and in response to indole-3-acetic acid and silver ions, Deduced amino acid sequences of OS-ERS1, OS-ERS2, OS-ETR2, OS-ETR3, and OS-ETR4 showed that they exhibited significant homology to the prokaryotic two-component signal transducer and a wide range of ethylene receptors in a variety of plant species
Prev Next